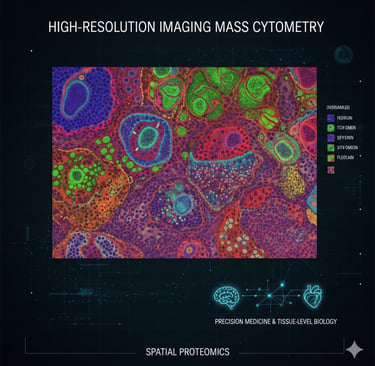
High-resolution imaging mass cytometry to map subcellular structures
High-resolution Imaging Mass Cytometry (HR-IMC) enables detailed mapping of protein expression directly in tissue. By combining oversampled imaging with computational deconvolution, it reveals subcellular structures while preserving high multiplexing. This approach advances spatial proteomics and opens new possibilities for precision medicine and tissue-level biology.
12/27/20251 min read
High-resolution Imaging Mass Cytometry (HR-IMC)
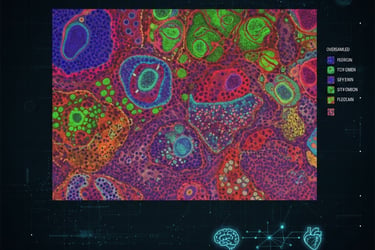
Imaging Mass Cytometry (IMC) is a powerful multiplexed imaging technique that uses metal-tagged antibodies and mass spectrometry to measure dozens of protein markers in situ. This new Nature Methods paper presents High-Resolution IMC (HR-IMC): a practical workflow that combines oversampled ablation with image deconvolution to push IMC from ~1 µm down to ~333–500 nm effective resolution, enabling visualization of subcellular structures such as nucleoli, mitochondrial networks and nuclear foci while keeping the high multiplexing advantage of IMC.
What the authors did
They collected IMC data with overlapping (oversampled) laser shots and applied deconvolution (e.g., Richardson–Lucy) to reconstruct higher-resolution images from the ion-count maps. This computational acquisition tweak improves apparent spatial resolution without changing the instrument hardware. The method was validated across sample types (tonsil, ovarian cancer models and others) and compared with classical IMC and aligned immunofluorescence (IF) images to confirm improved subcellular localization and marker separation.
Key findings & why they matter
Subcellular detail: HR-IMC resolves organelle-level features (mitochondrial patches, nucleoli) and
distinct nuclear/membrane marker patterns that classical IMC (1 µm) blurs.
Better segmentation & phenotyping: Improved resolution reduces mixed/ambiguous cell masks
(for example, previously inseparable B-T doublets), leading to more accurate cell segmentation and
pixel-level clustering for downstream analysis.
Multiplex + resolution: HR-IMC keeps IMC’s strength (high multiplex, large dynamic range, no spectral overlap)
while making in-situ proteomic maps more comparable to light microscopy in spatial detail — valuable for
tumor biology, spatial proteomics and spatially resolved functional studies.
Takeaway for readers / researchers
HR-IMC is an accessible technical advance: by changing acquisition step size and using deconvolution, labs with standard IMC instruments can generate higher-resolution proteomic images that reveal subcellular organization and improve cell-level analyses. This bridges the gap between highly multiplexed spatial proteomics and the subcellular detail researchers expect from optical methods — opening new possibilities for studying organelle dynamics, subcellular biomarker localization and treatment responses directly in tissue.
https://www.nature.com/articles/s41592-025-02889-8

Follow
Contact
info@microlab.ink
© Microlab-copyright 2026

